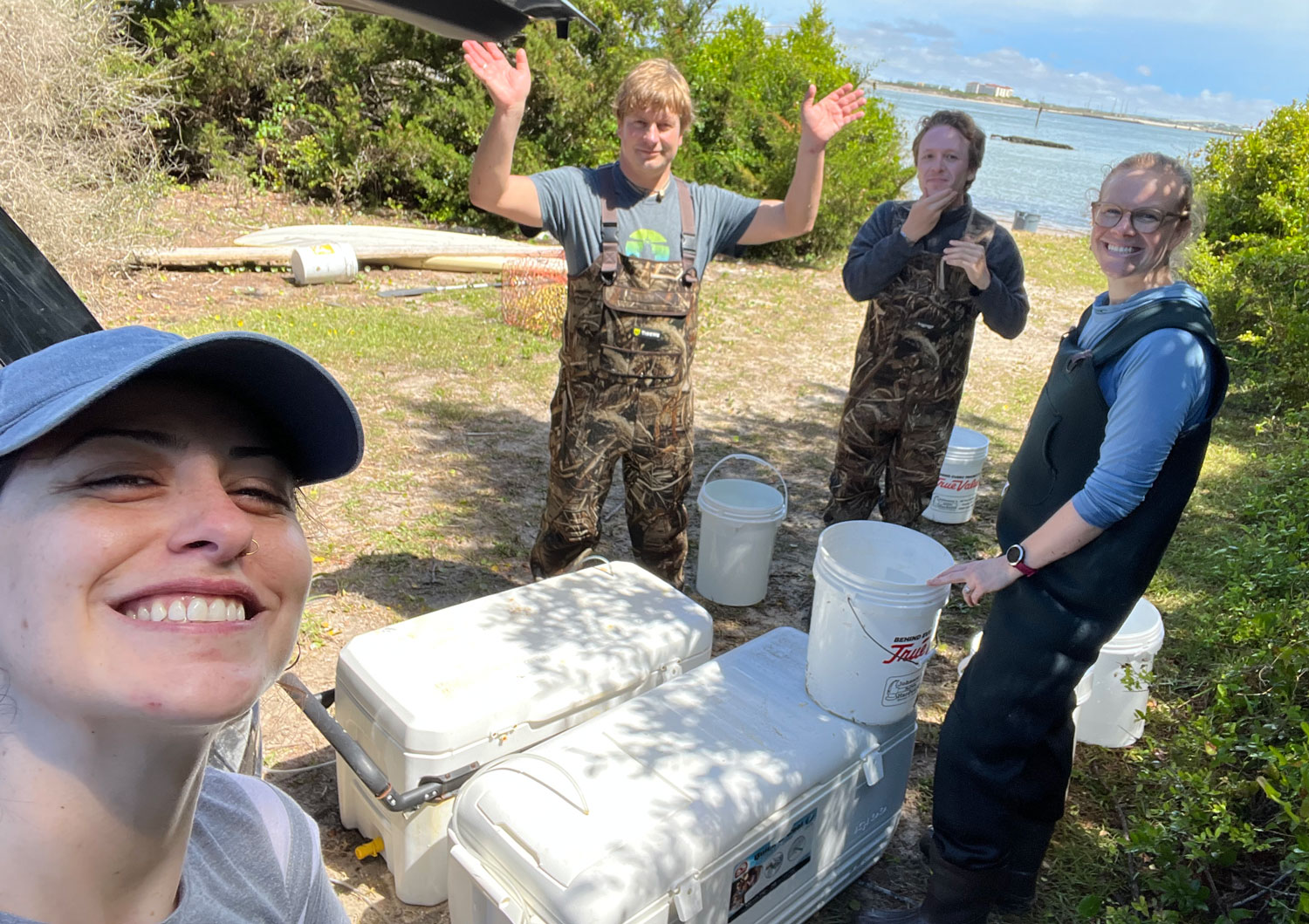
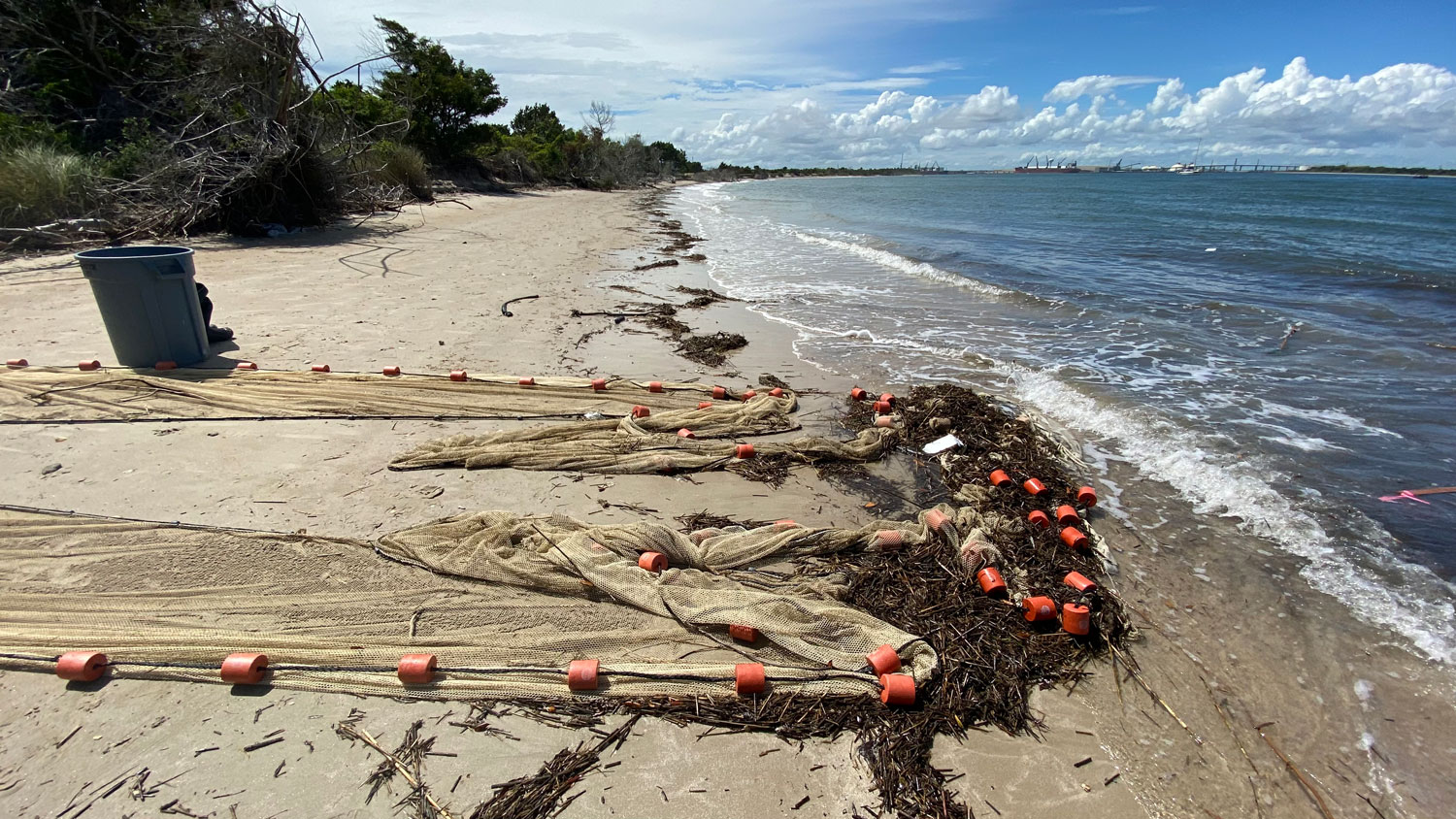
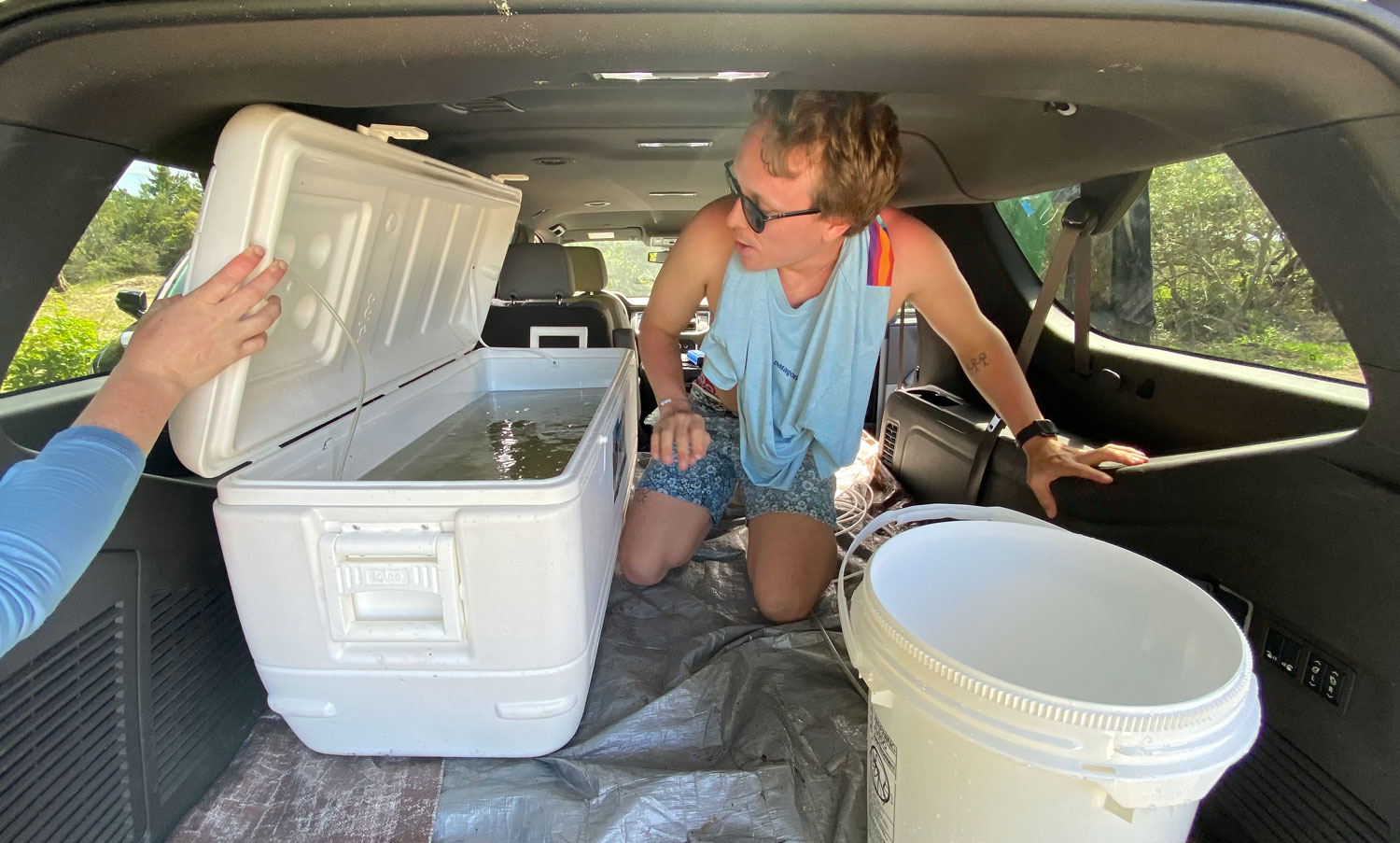
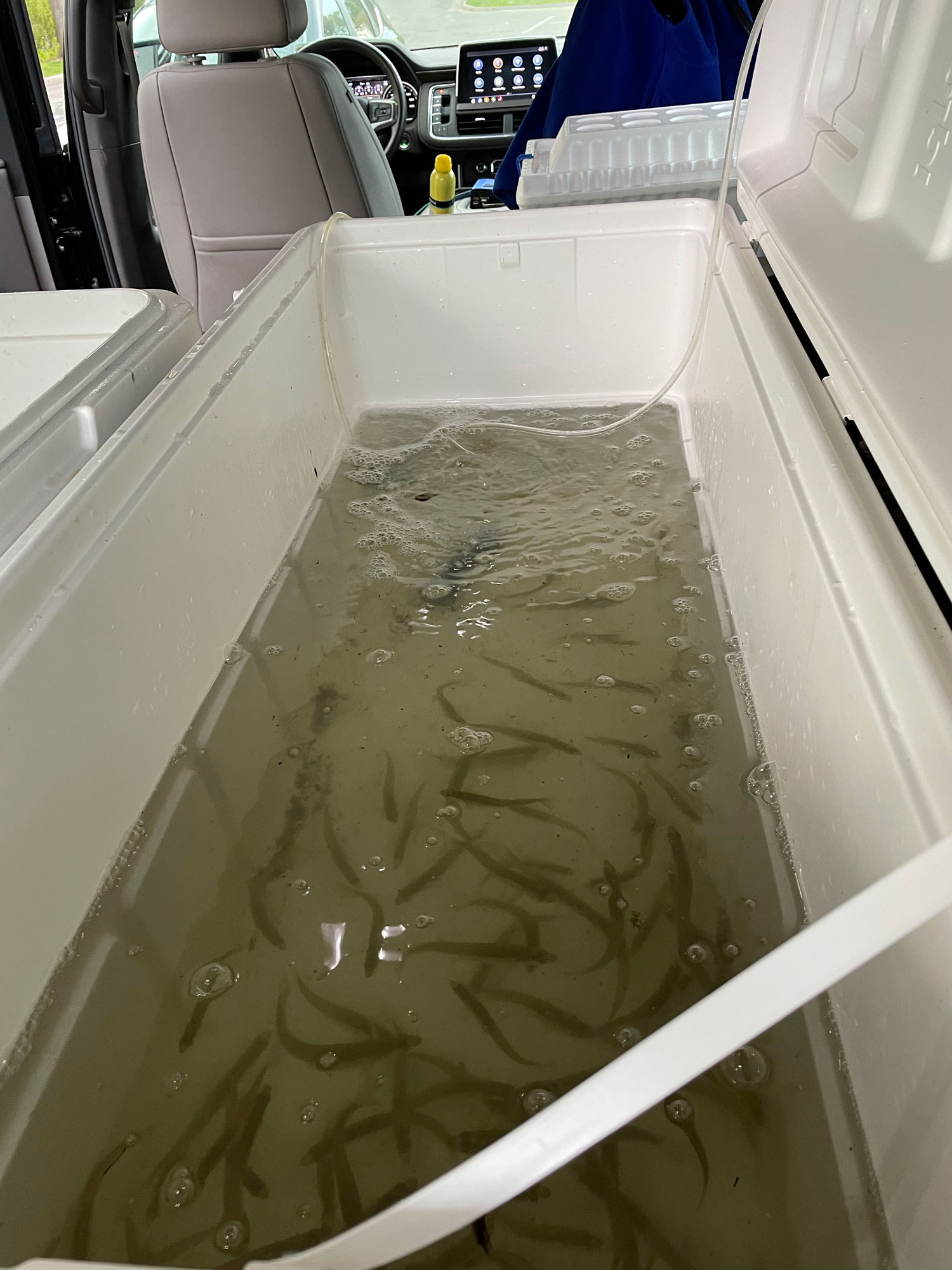
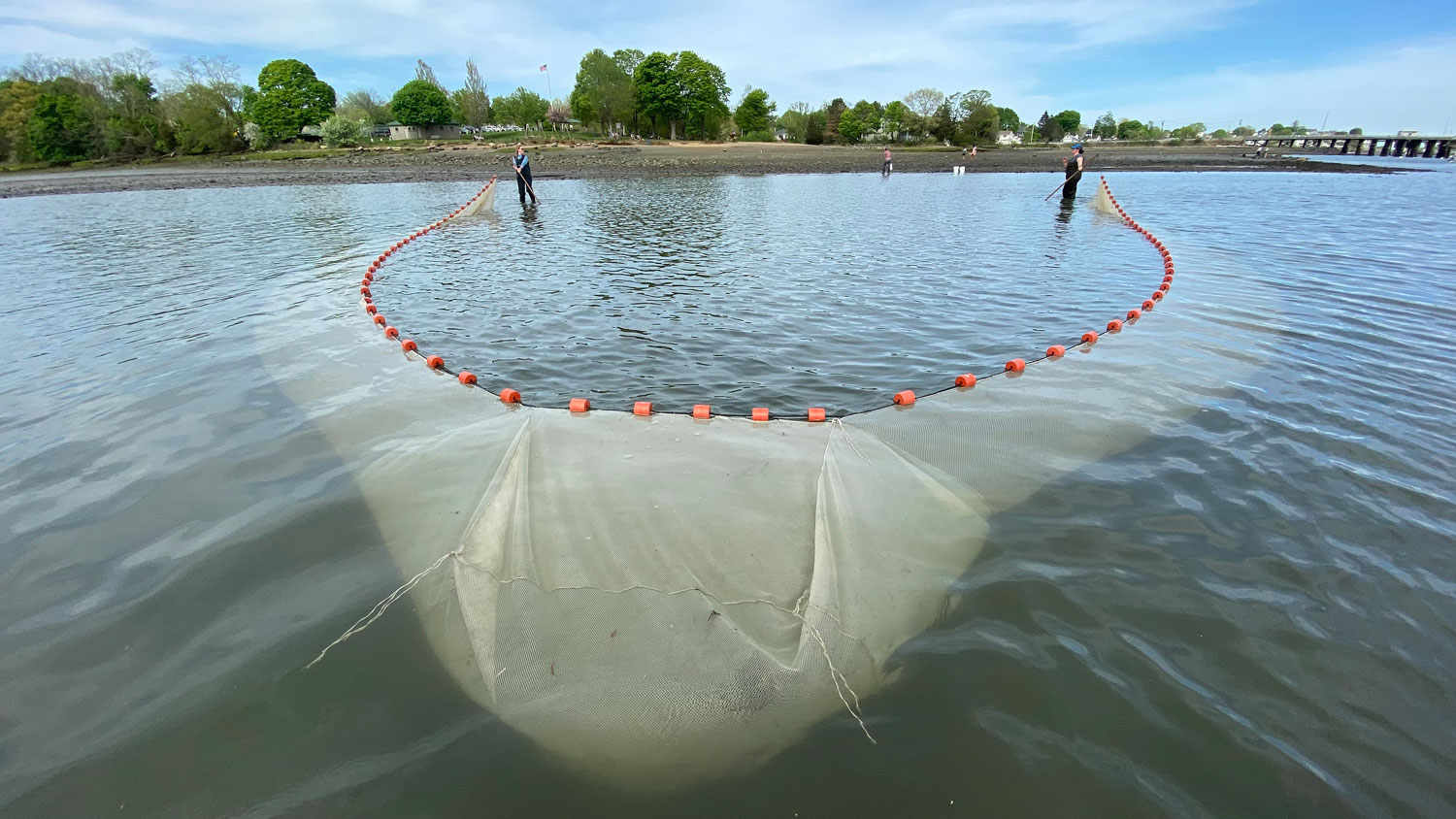
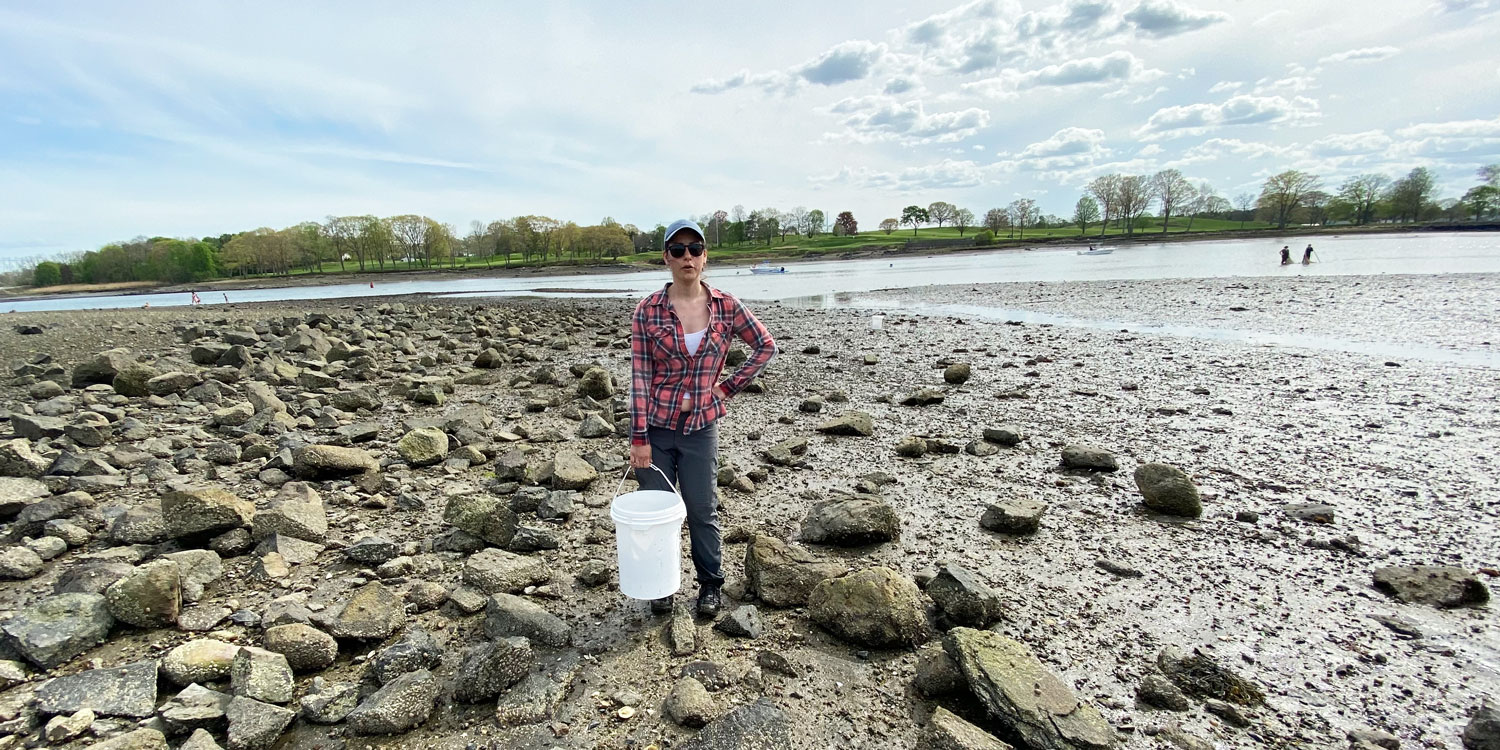
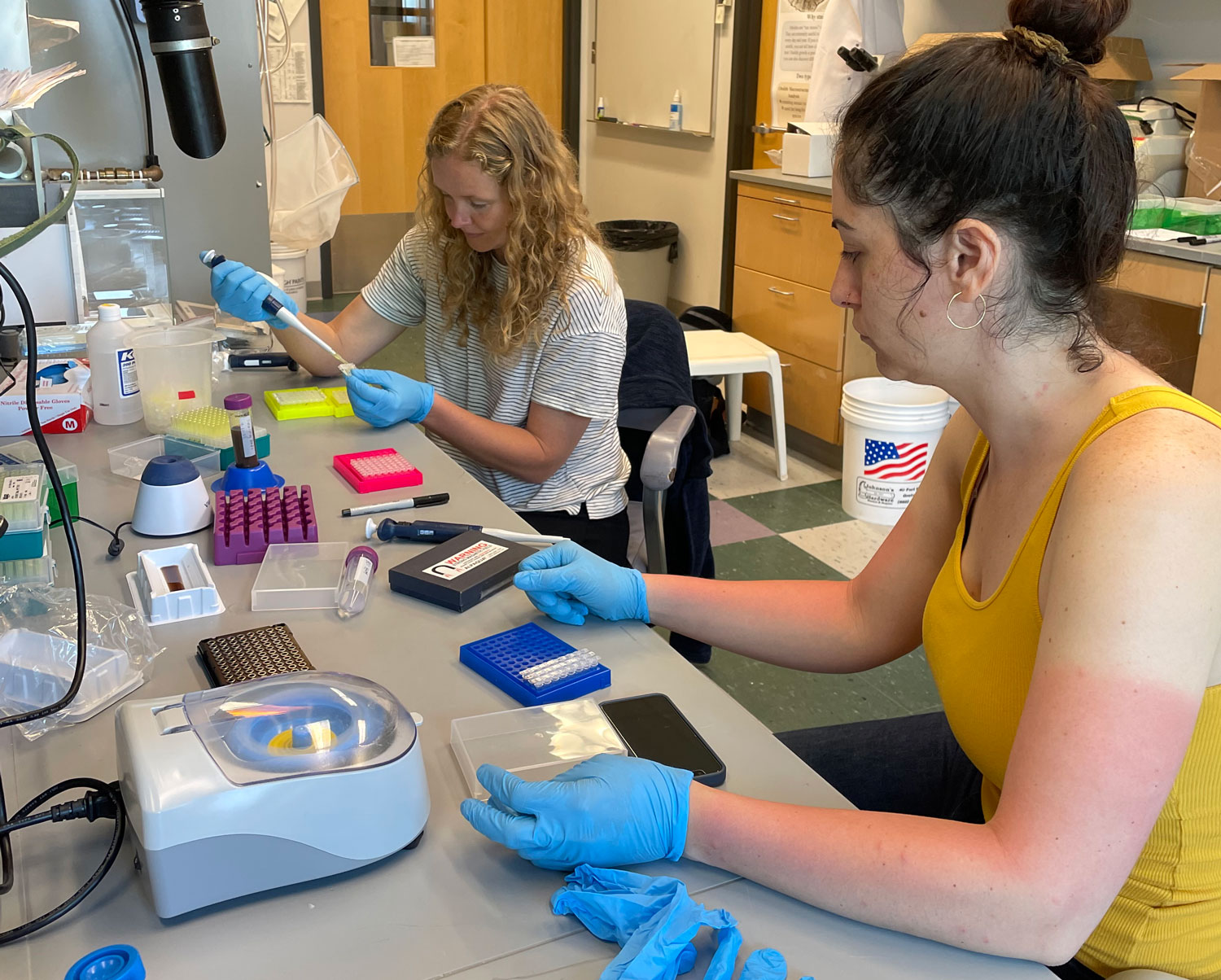
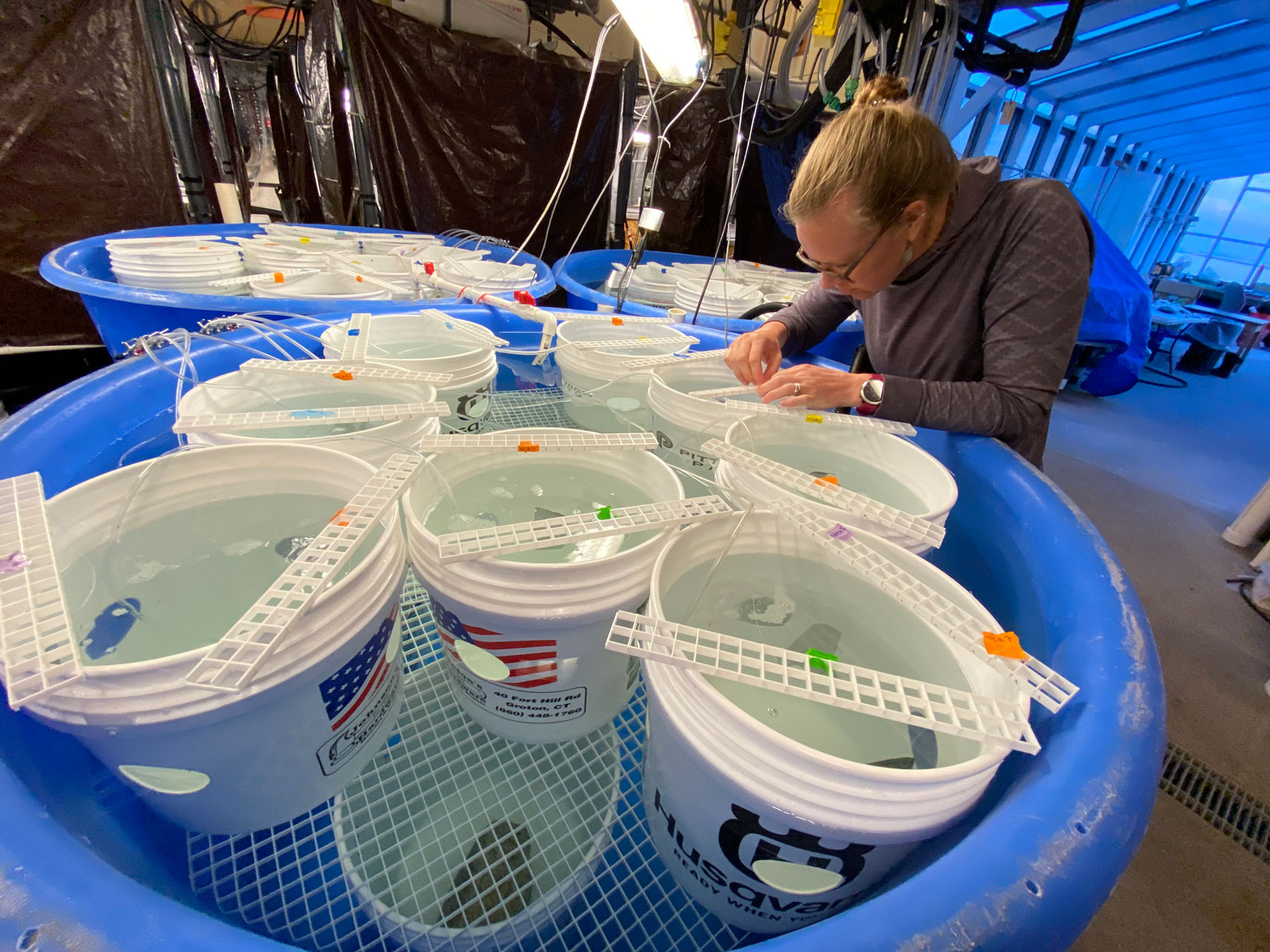
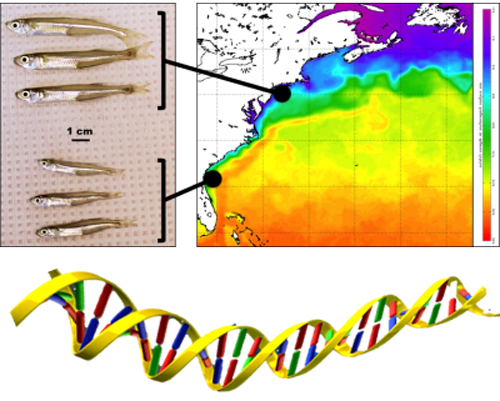
25. September 2025. I have been asked by quite a few people (on both hemispheres), why I had to go again to Chile, to the same place, the same marine station, to repeat the same experiment from two years ago? Did the first fail somehow? No, not at all. The first experiment yielded really intriguing data suggesting similar local adaptation patterns in southern compared to northern hemisphere silversides.
But the catch is that one year could be a coincidence. To make statistically robust inferences about the nature of local adaptation in Chilean silversides, good science simply demands another, a second independent data set of observations. Even more so, because (i) the adaptation strength here is likely subtle, and (ii) some treatments and populations were indeed less successful the first time around.
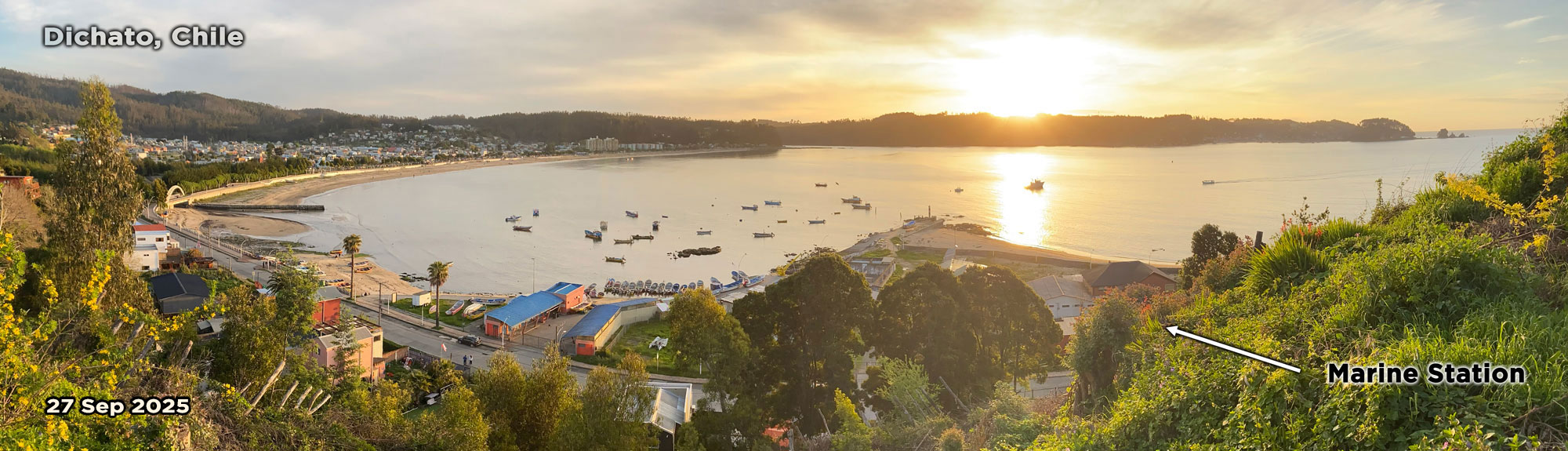
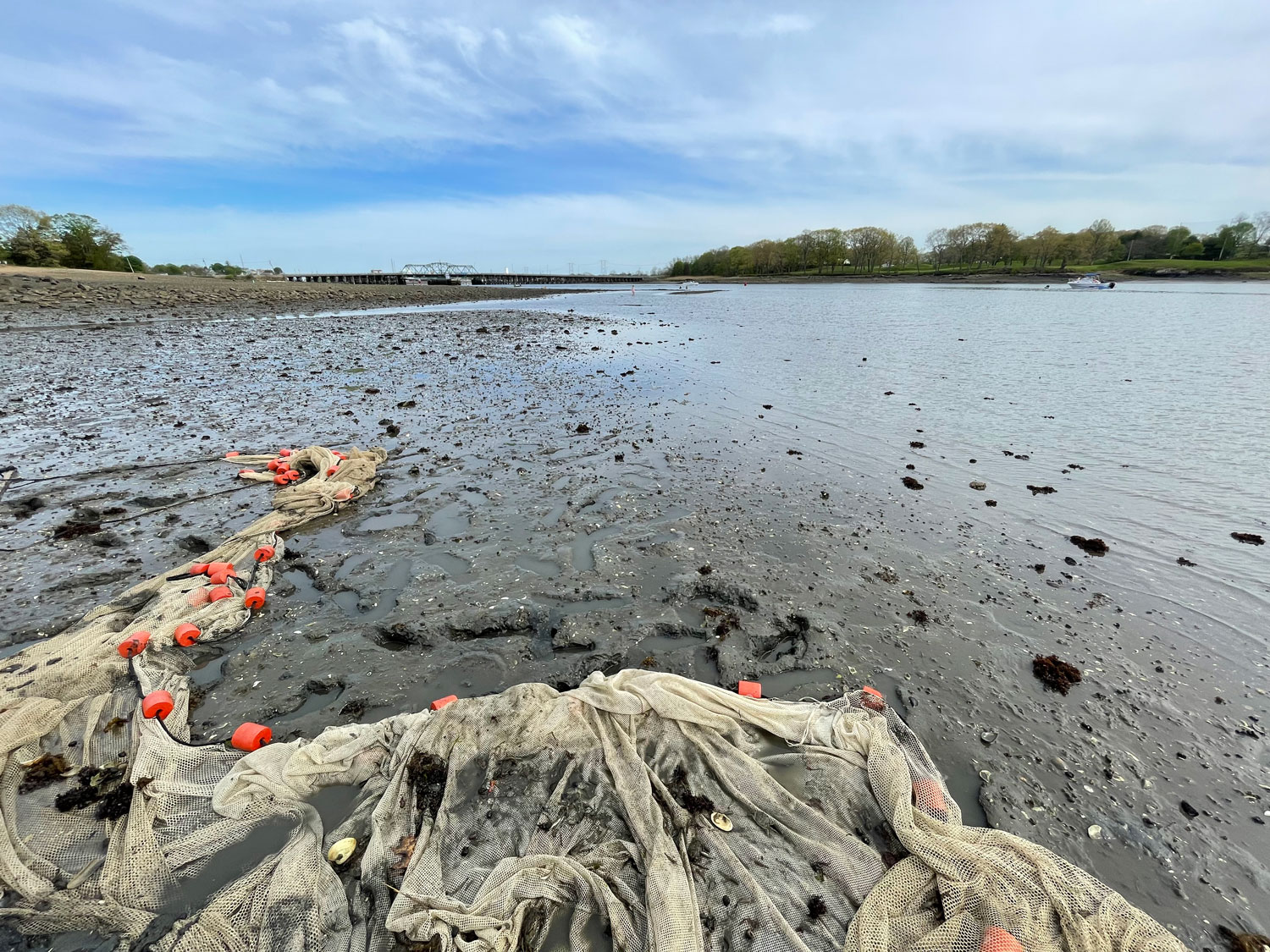
And so I'm here again. The place where I spent 6 months during my sabbatical feels wonderfully familiar - despite being thousands of miles away from home. The setting September sun pours gold over the halfmoon-shaped Coliumo Bay. Spring is in the air here, but the little beach town is still mostly void of the summer crowds. I wander through the streets, recognize the stray dogs, and many of the people in this little village say they remember me and my family from two years ago.
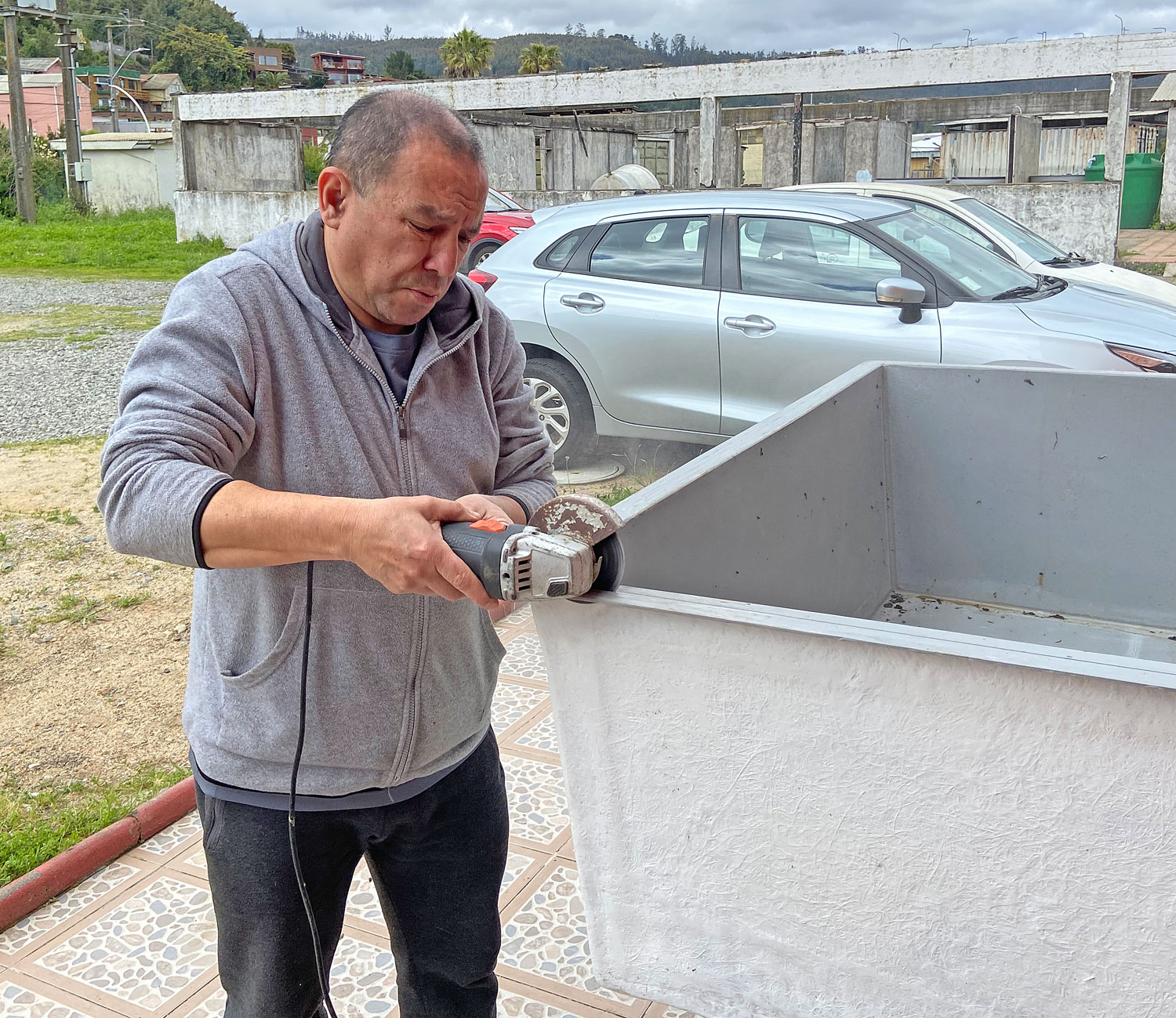
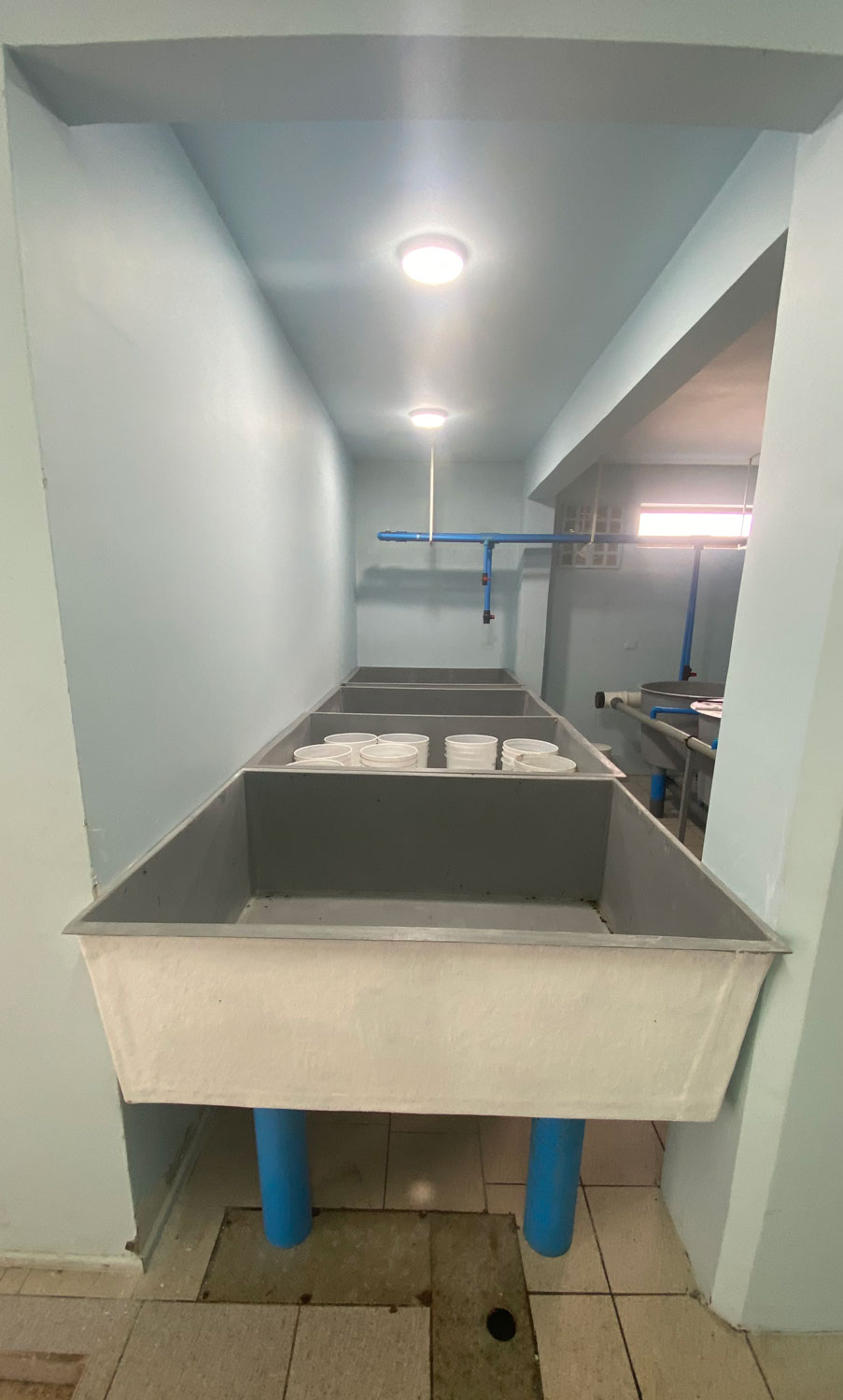
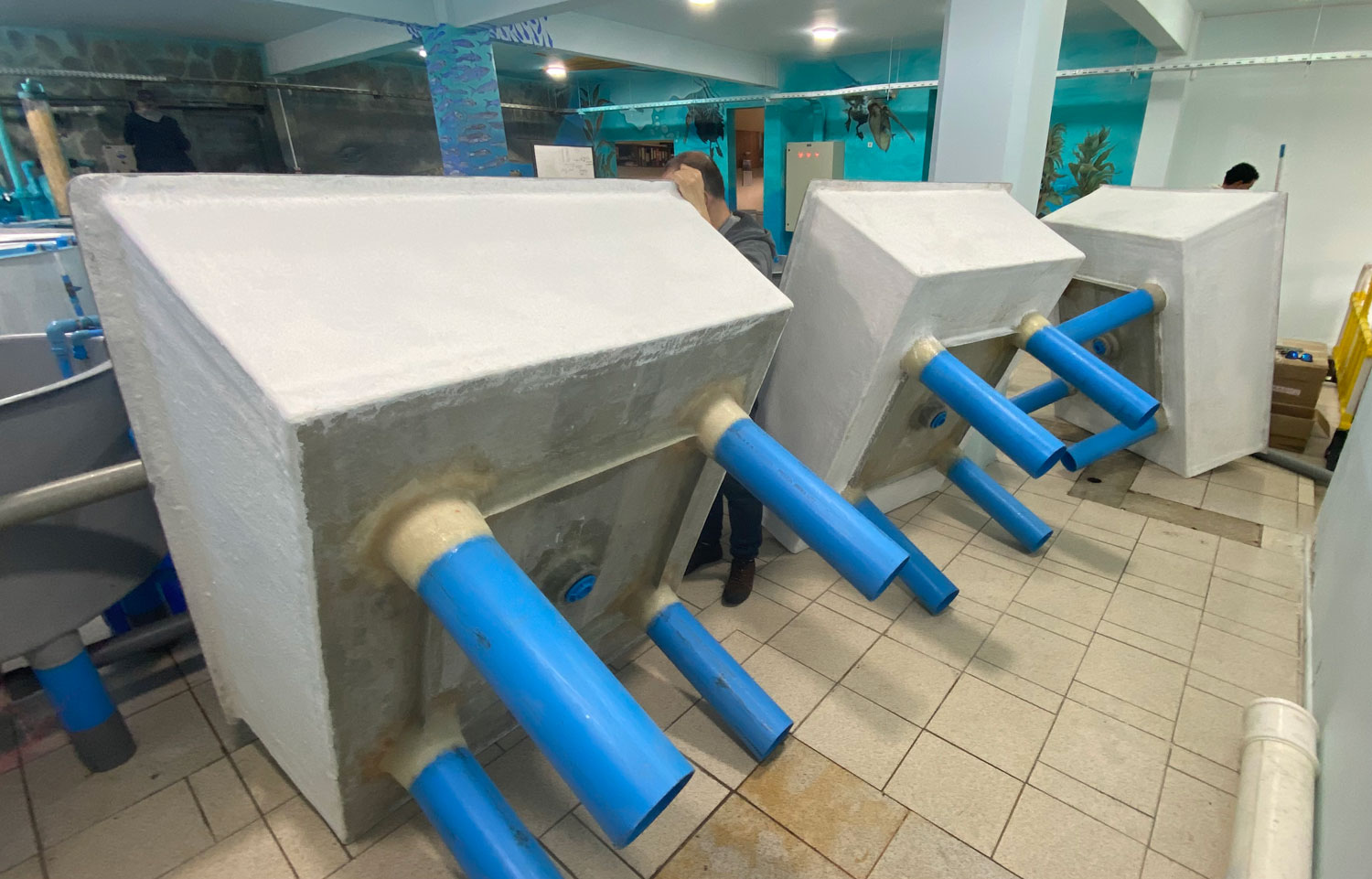
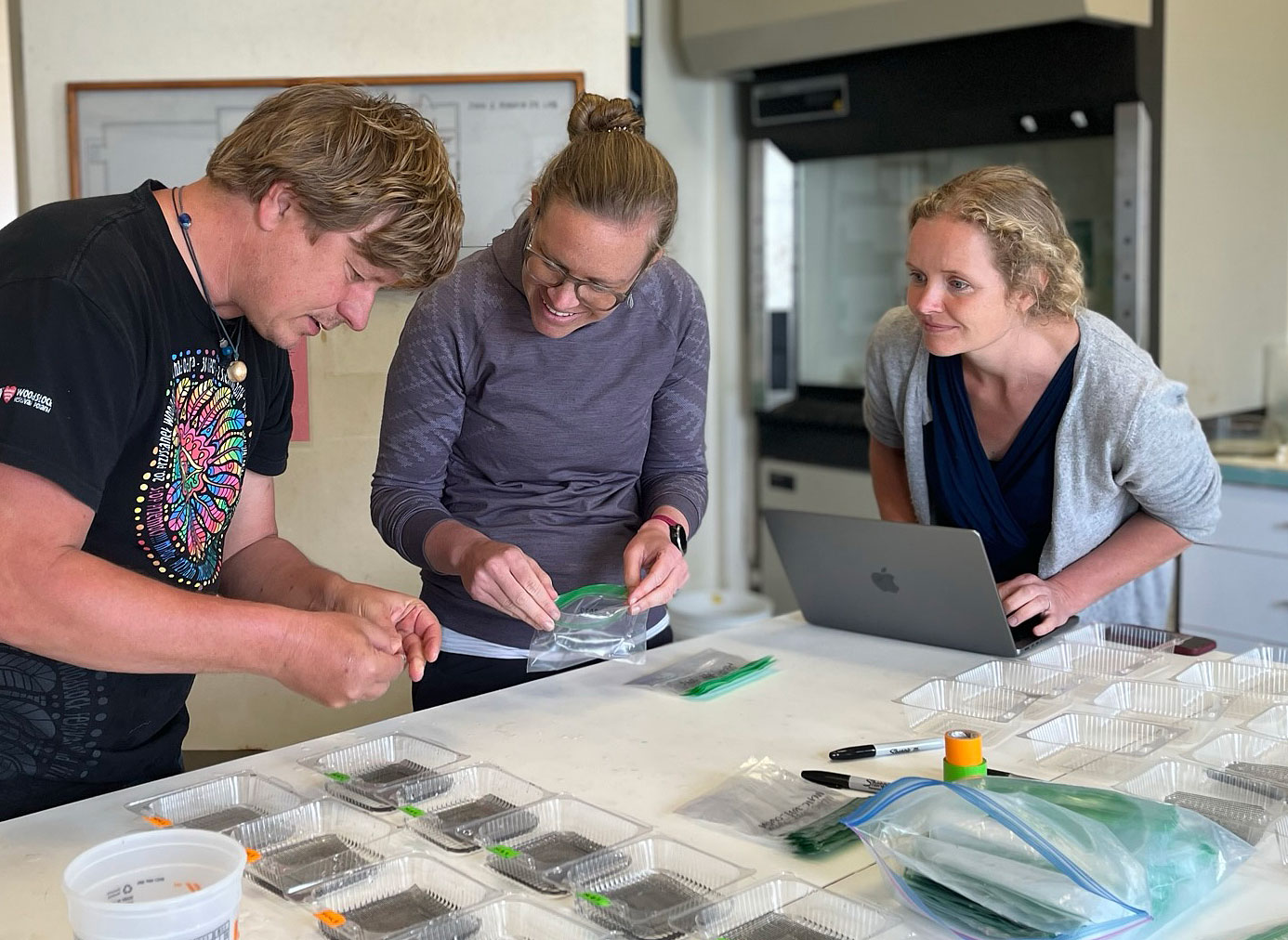
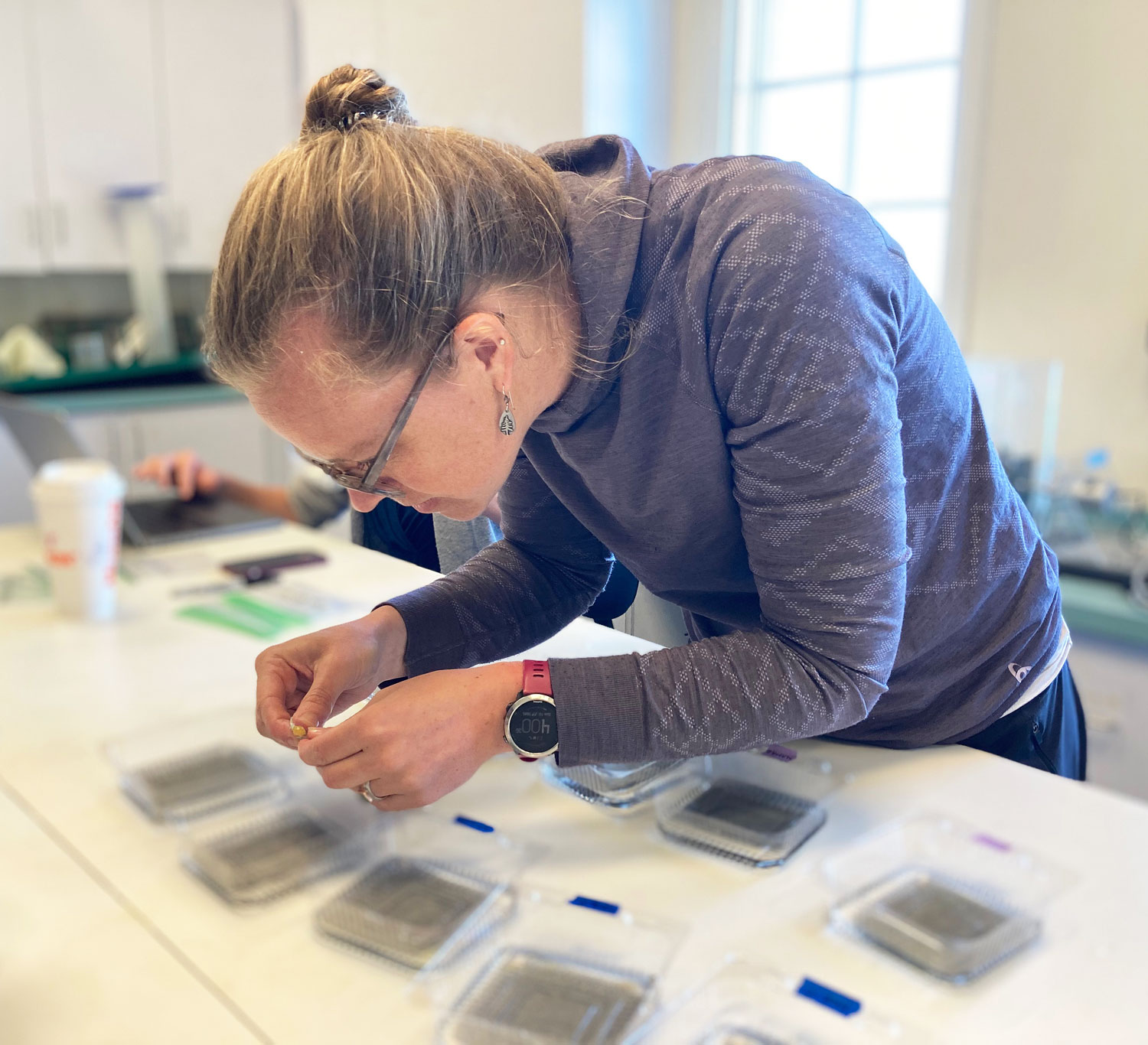
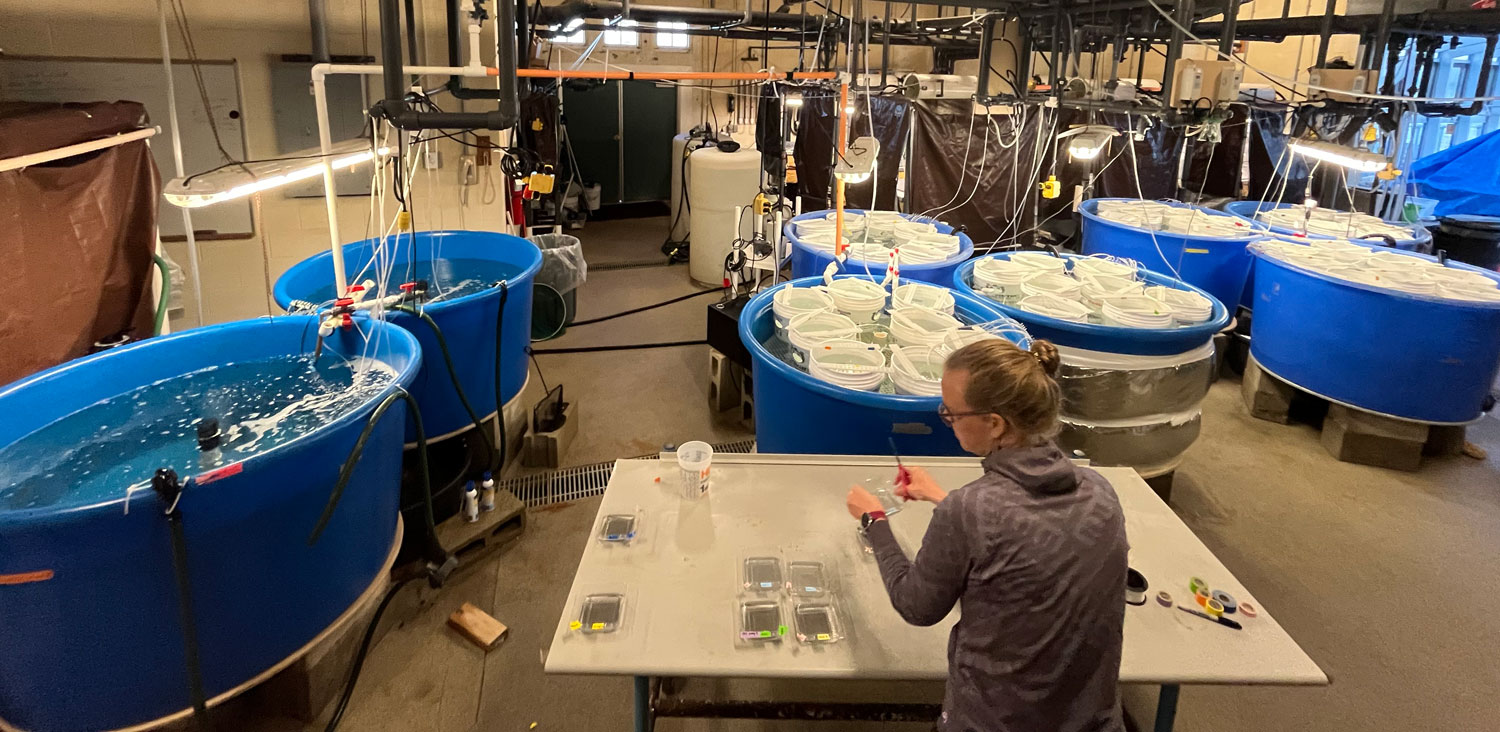
In the marine station, the experiment is now being set up in a different location. Inside the aquarium, which is climate-controlled and therefore more suitable. It took us two years to gain permission to move in, and I had to have custom tanks made to set it all up. Will it all be worth it?
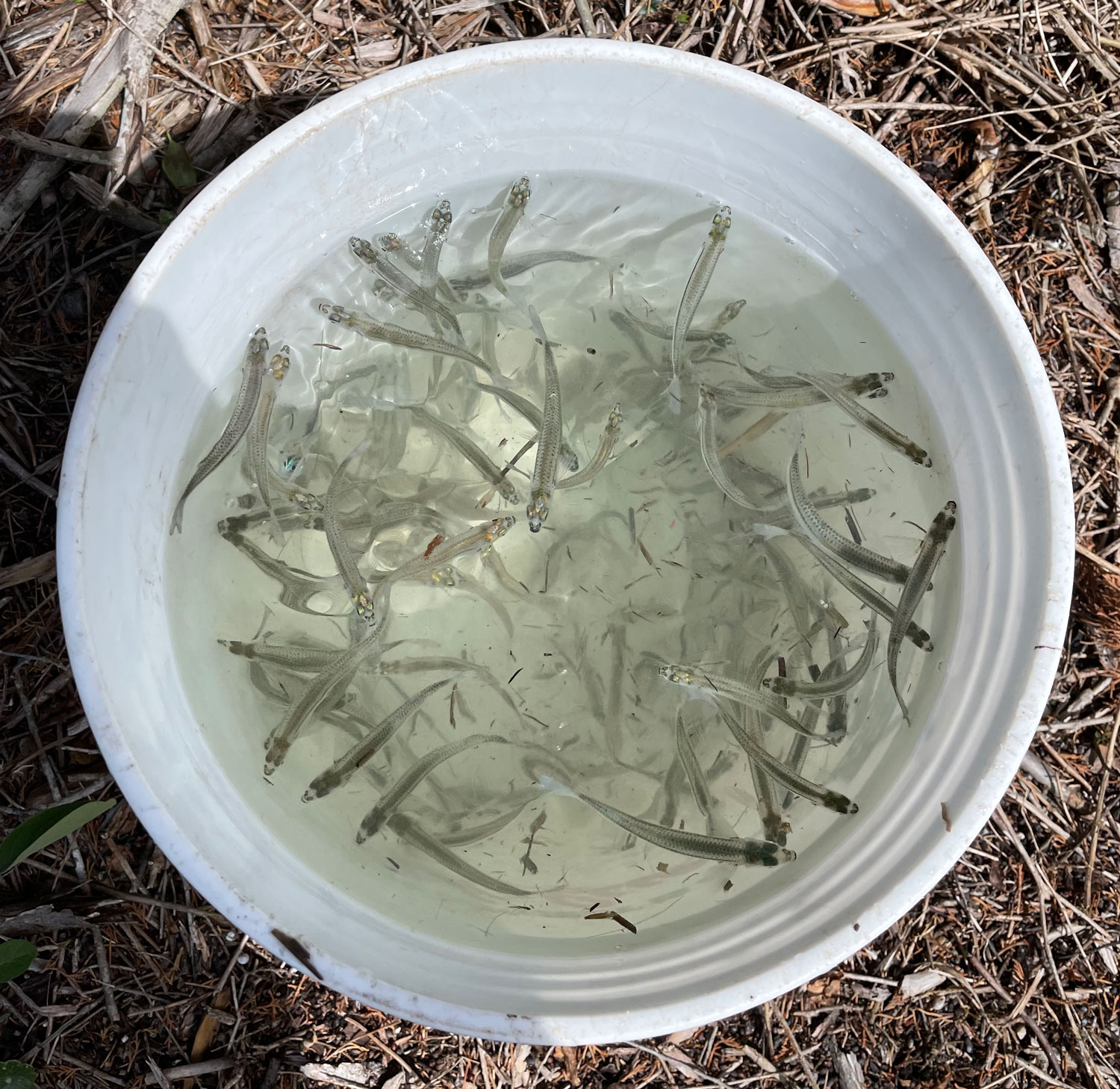
Now time is really tight, the setup needs to be put together in just a few days, because in the low latitudes of Chile's north, the spawning season of pejerrey has already begun. Fingers crossed.