By Emma Siegfried.
In 2019, the Evolutionary Fish Ecology lab at the University of Connecticut led by Dr. Hannes Baumann published their first paper investigating the effects of ocean acidification and warming on Ammodytes dubius. They found that levels of ocean acidification predicted for the year 2300 significantly decrease hatching success of embryos. The finding has begged the question if congener species are similarly susceptible – indicating the potential evolutionary conservation of this trait.
I began my PhD with the goal of doing similar experiments to other closely related species. One location where this would be possible is at the Institute of Marine Research (IMR), where a few scientists, notably Prescilla Perrichon, the leader of the Norwegian AQUASERV programme, have been spawning and raising Ammodytes marinus for a few years now. I applied to the AQUASERV transnational access program from the European Union and was fortunate to receive funding.
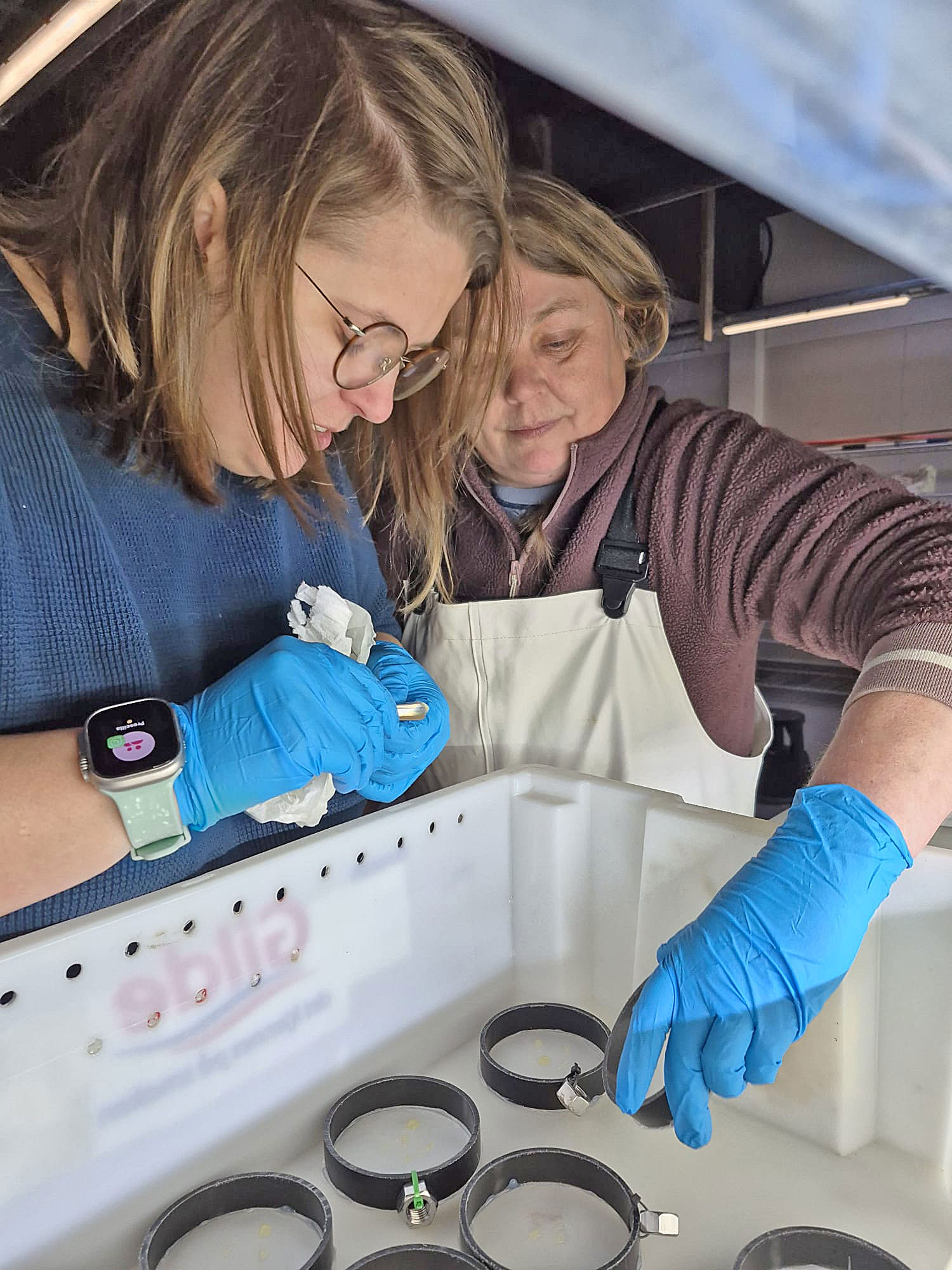
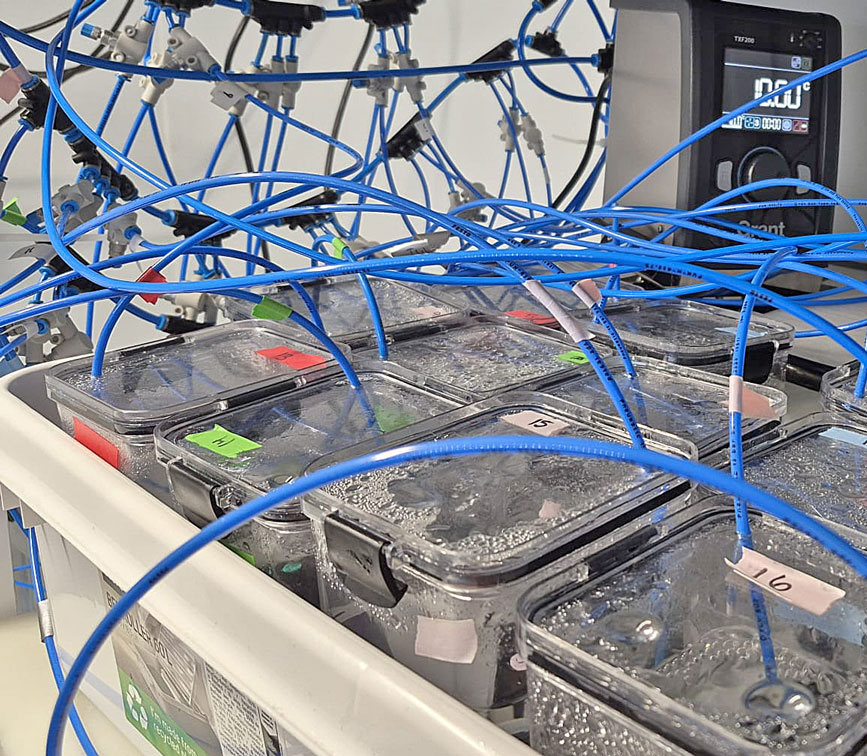
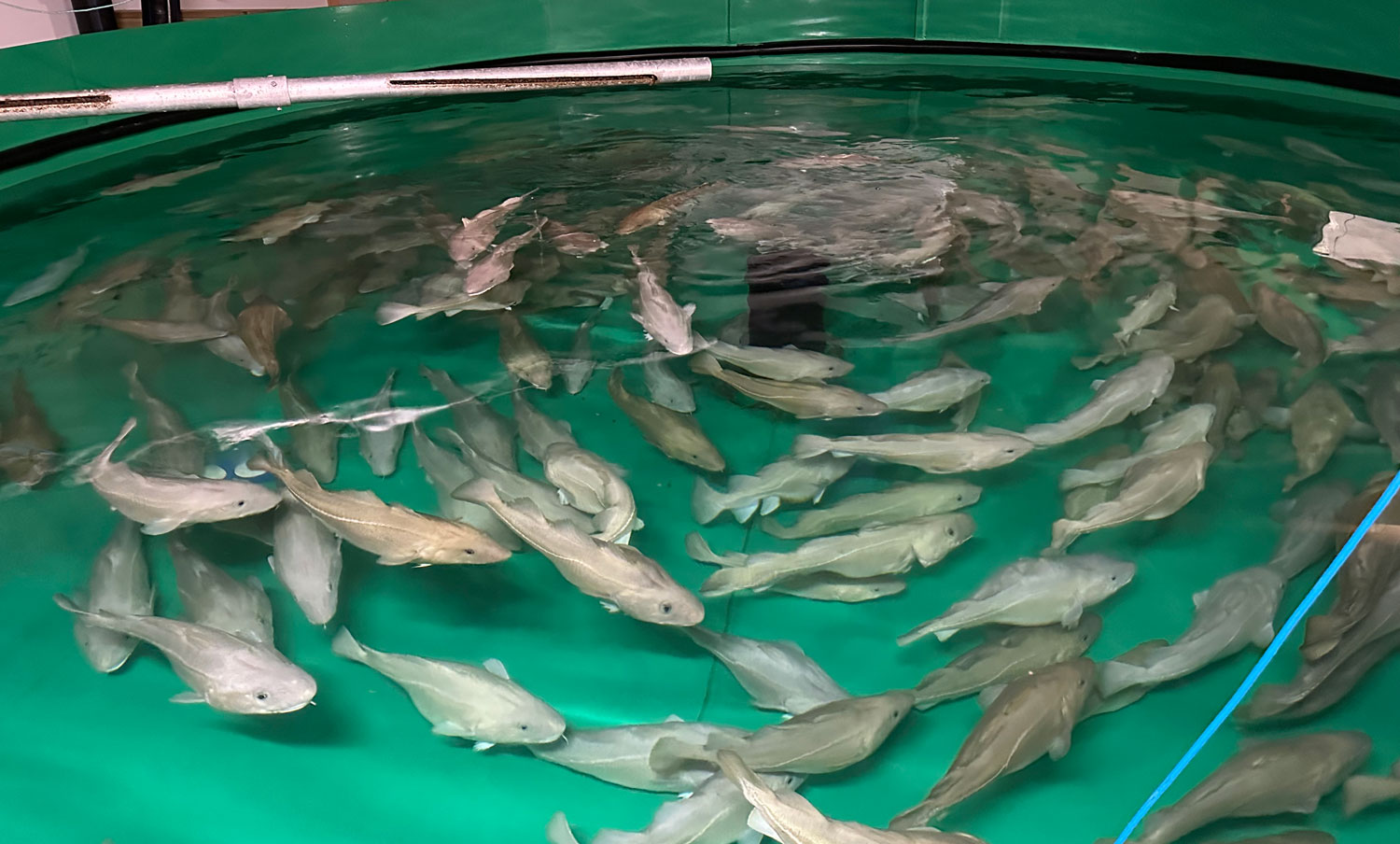
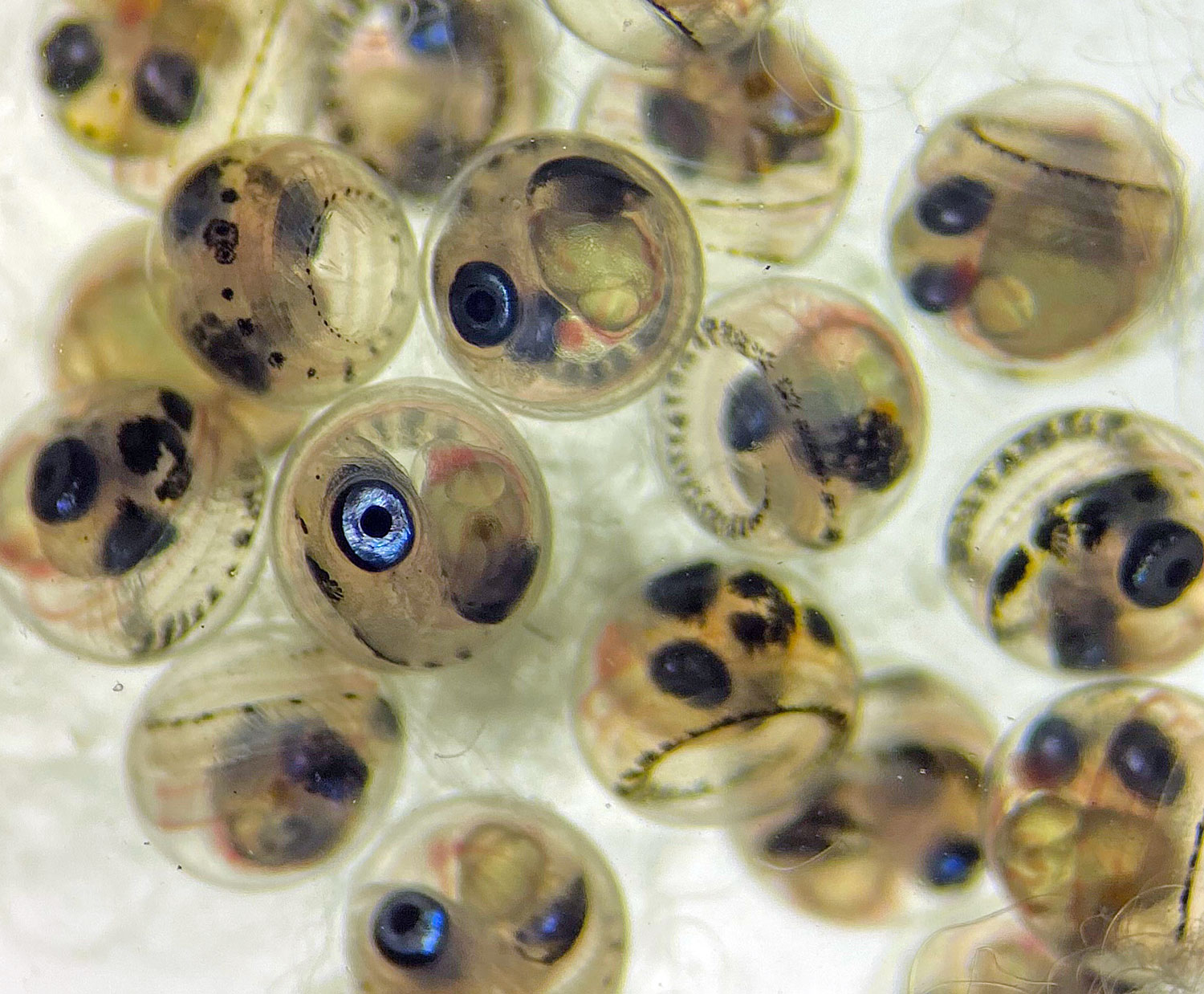
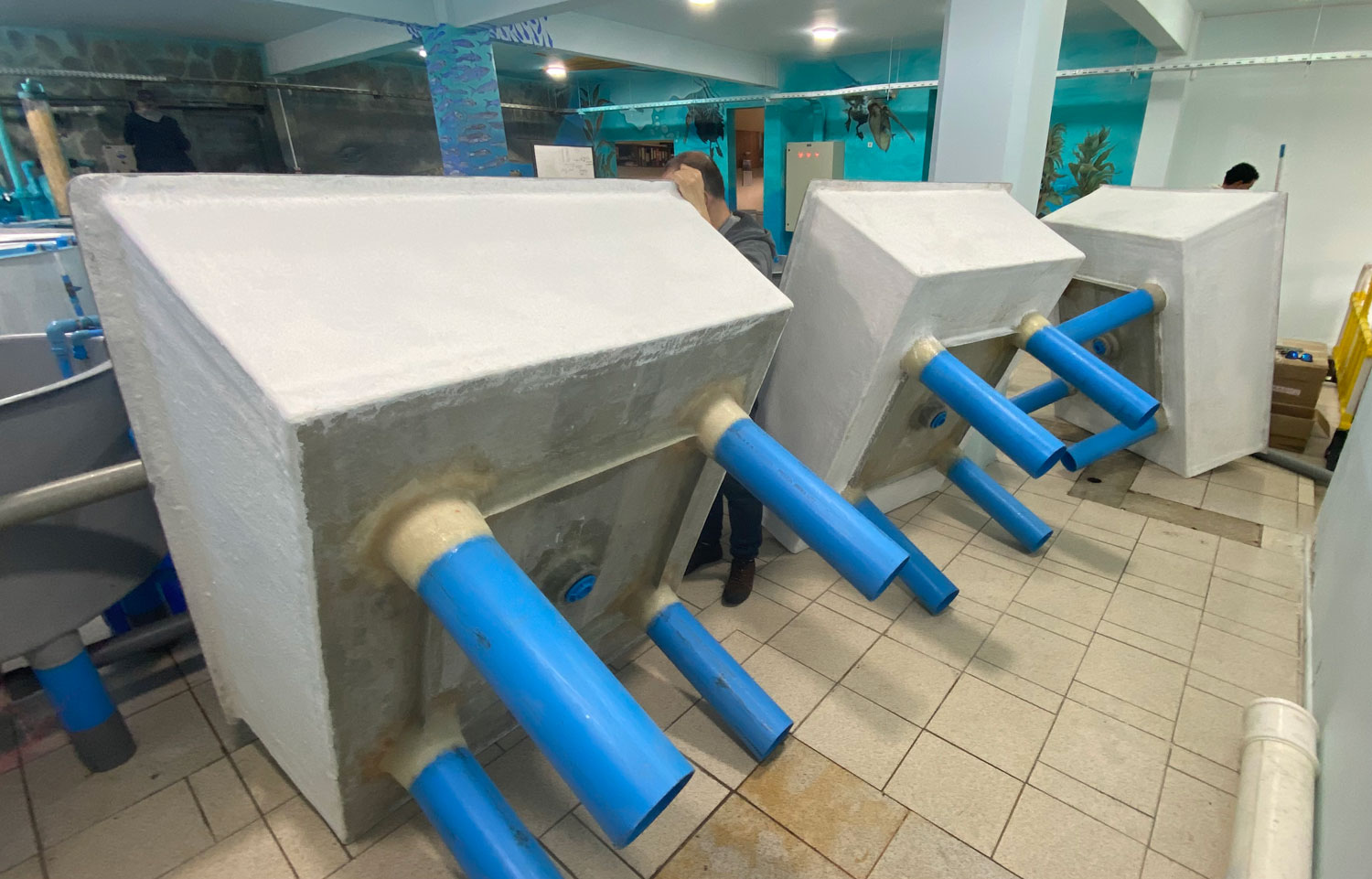
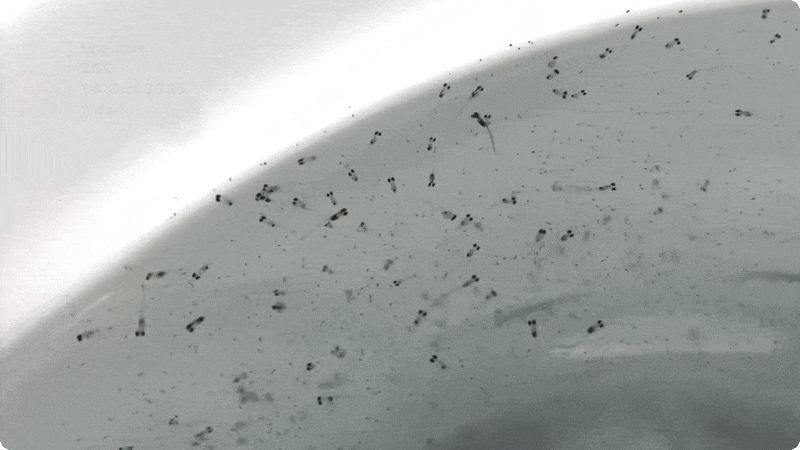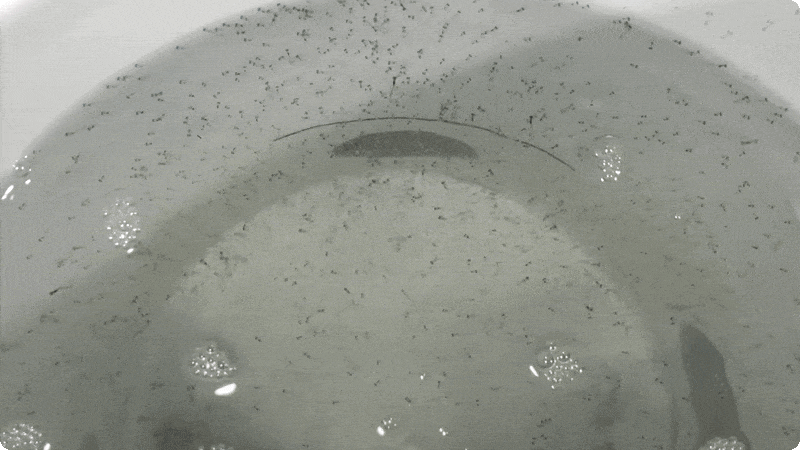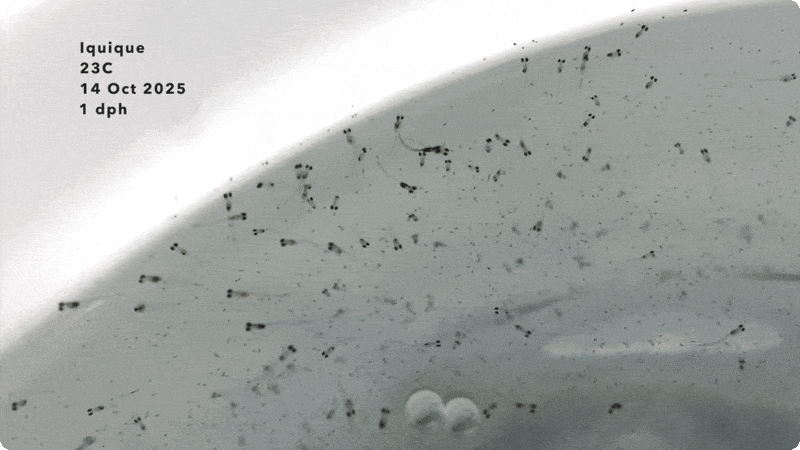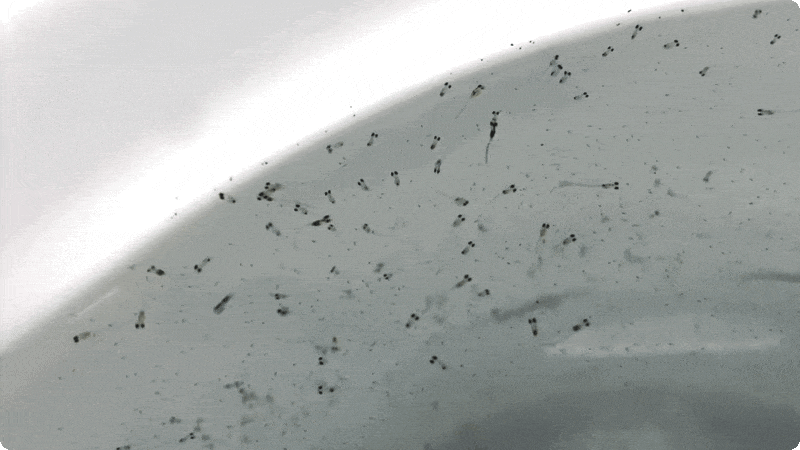
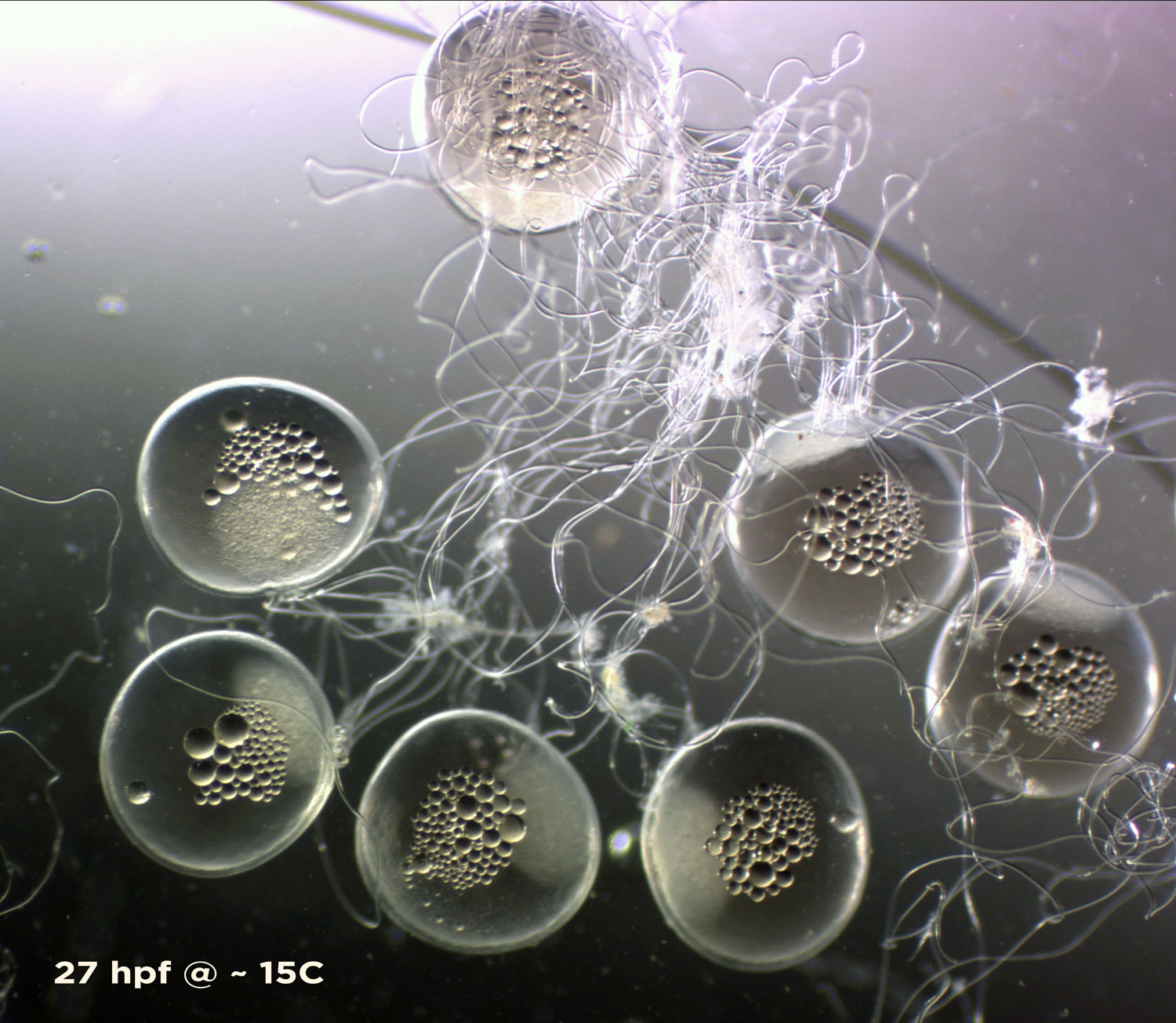
I arrived at the Austevoll Research Station at IMR in December 2025 unsure of what my time in Norway would bring. I had attempted similar experiments with Ammodytes americanus the previous winter and was not successful. In mid-January, the adult A. marinus that Reidun Bjelland had gone out and caught before I arrived began spawning. The fertilized embryos were placed in a rearing system, created with the help of Helen and Sam Rastrick, which exposed them to a combination of two different temperature and four carbon dioxide levels.
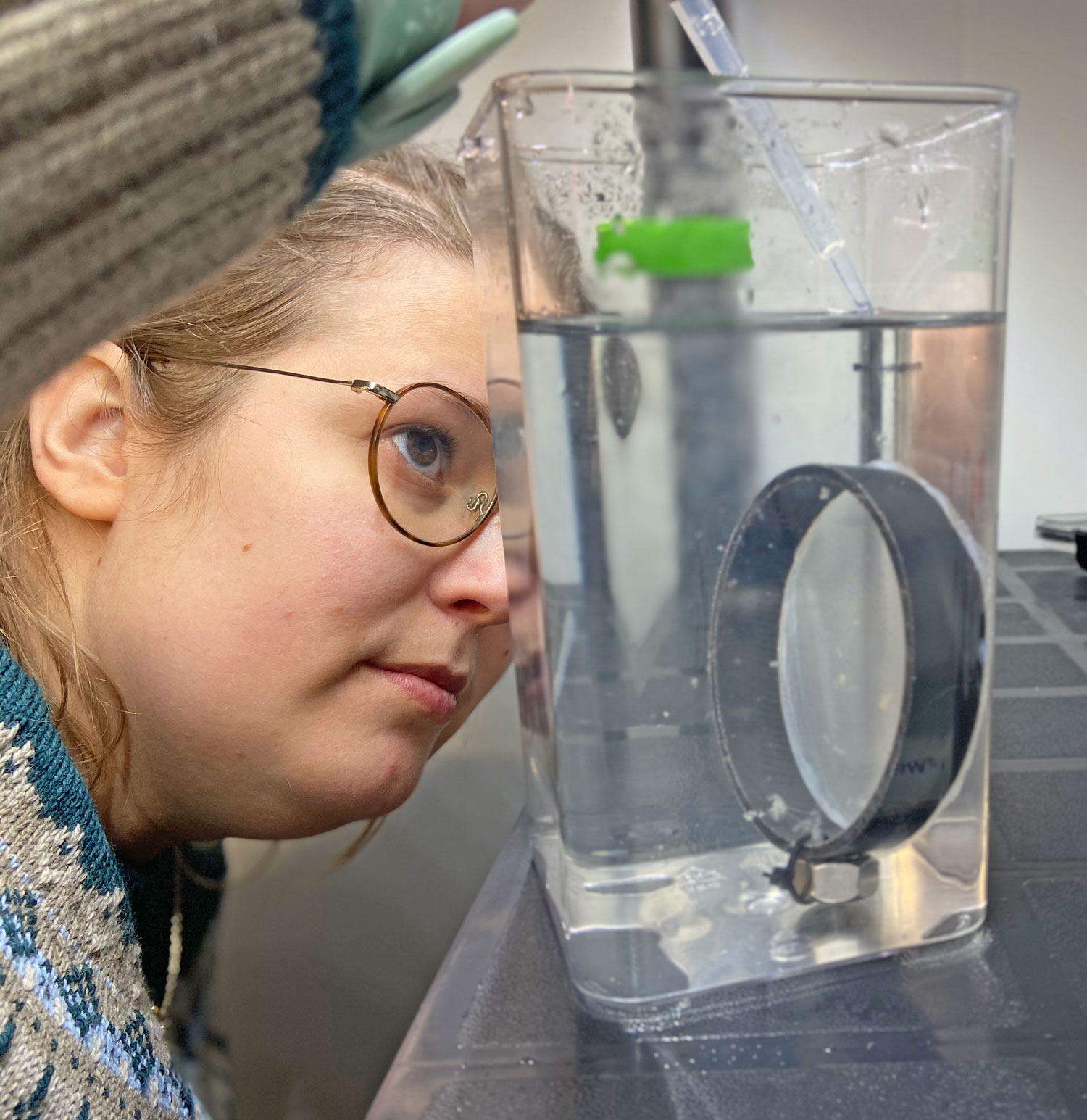
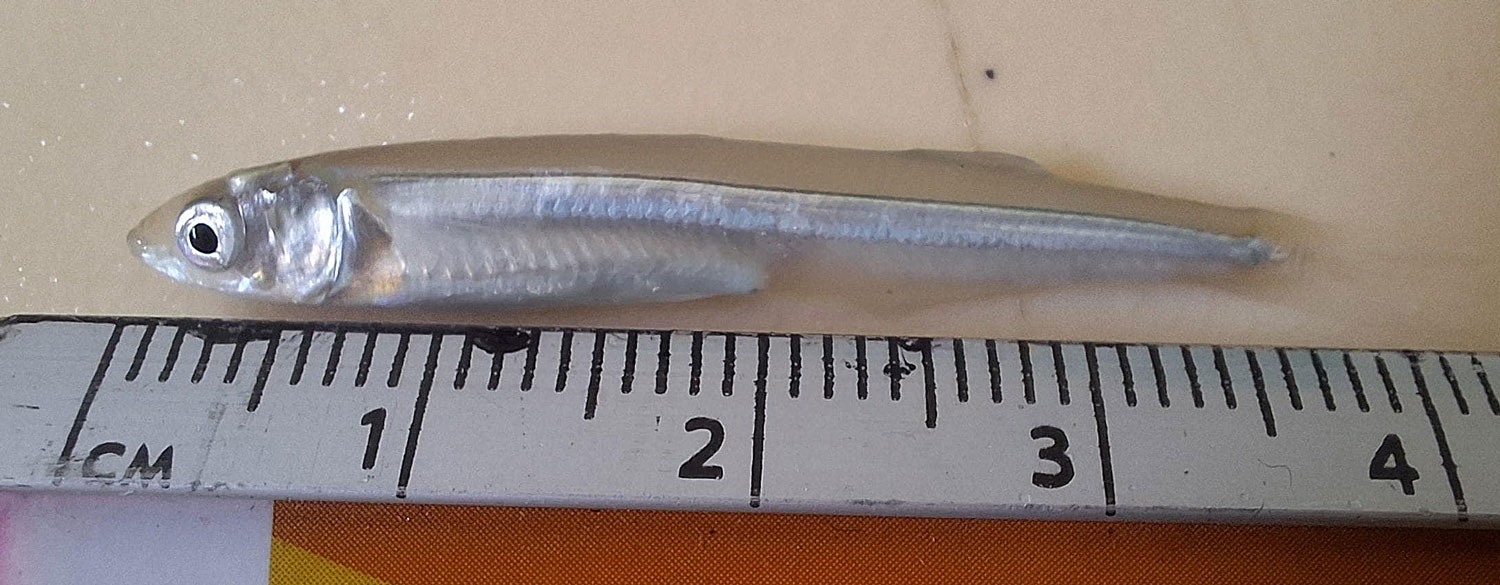
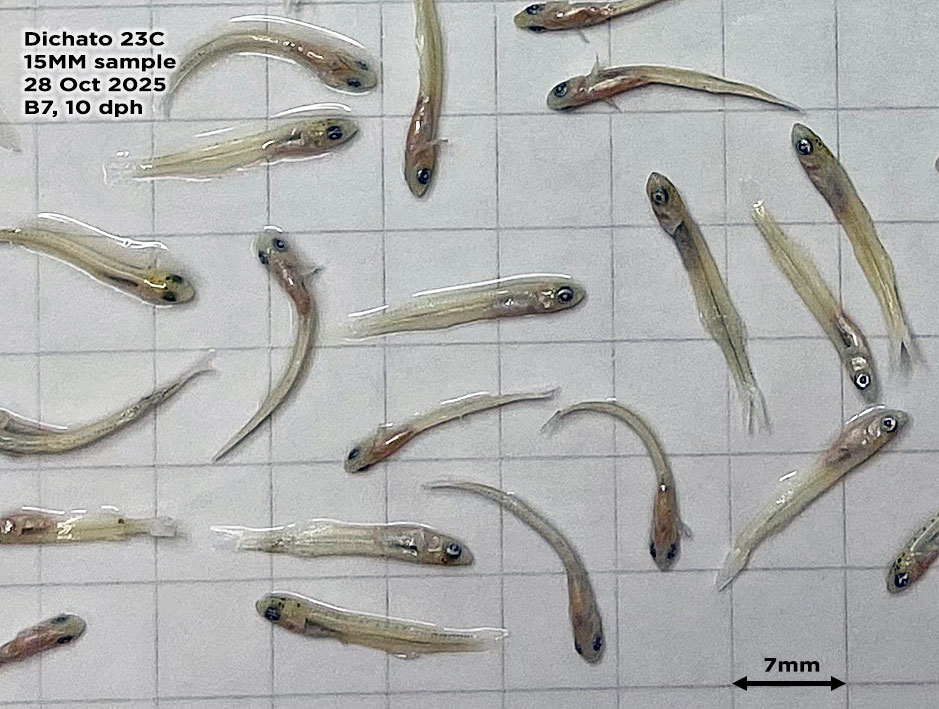
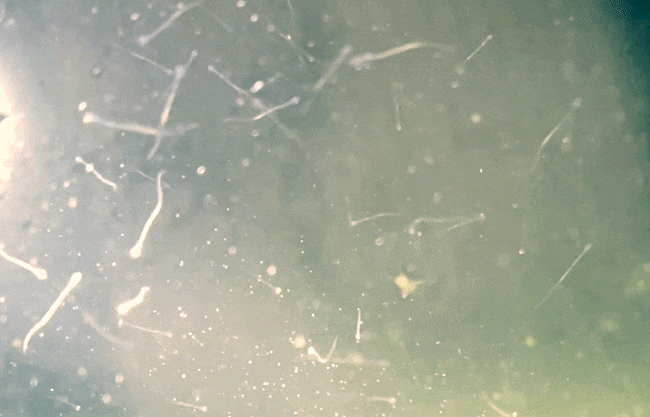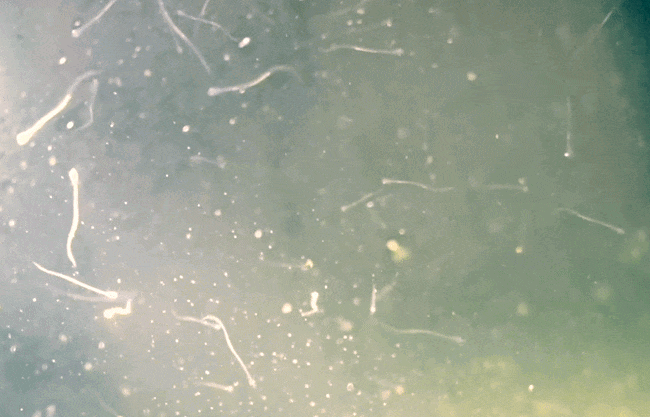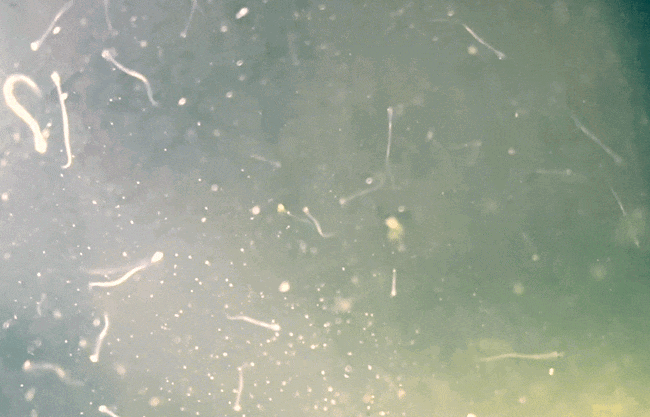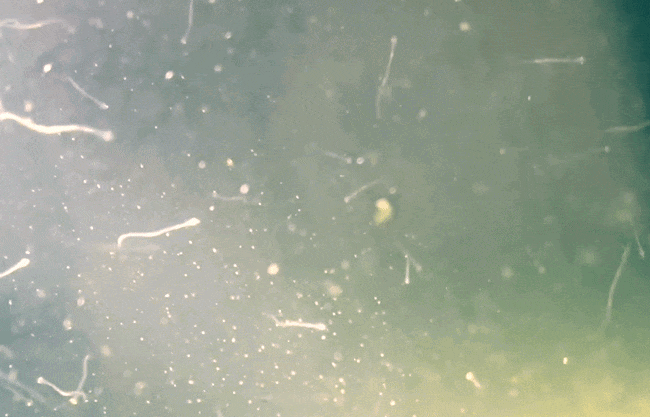
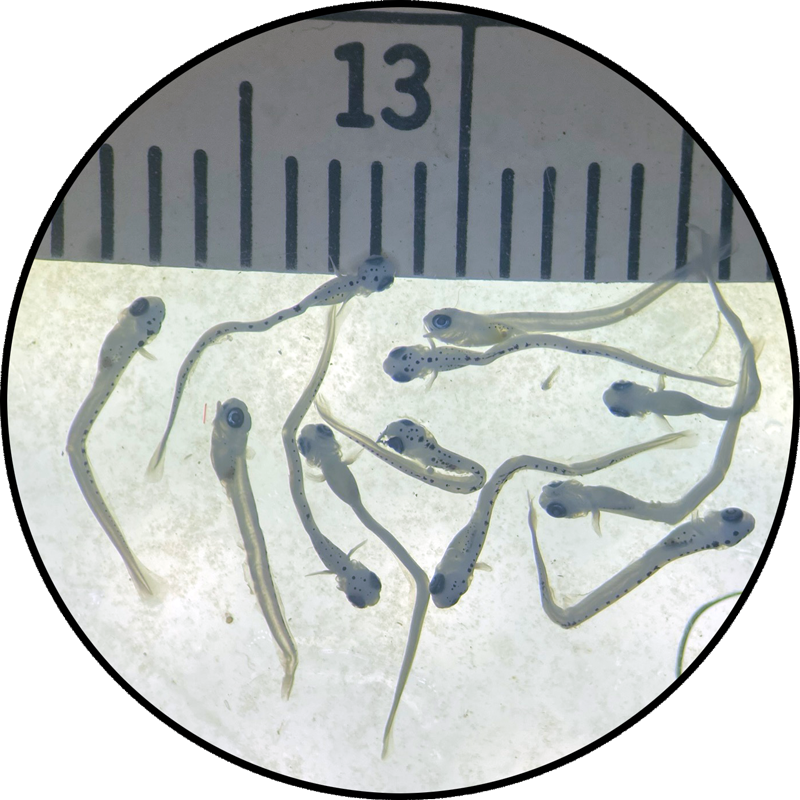
So far, we have collected data on the number of individuals hatched and how many embryos were in each individual tank initially. These values will be used to analyze hatching success and compare the impacts of ocean acidification and warming on A. marinus to the effects on A. dubius. In addition, we have photos of larvae that we can use to measure length, body area, and energy reserves. Finally, videos of the larval heart beating were taken for measurements of heart rate. In time, all this data will be analyzed and published.
I’m continuously grateful for the time I was able to spend at the Austevoll research station. It was truly the experience of a lifetime. I came out of my time in Norway with more knowledge and experience than I even thought possible and some new friends to boot. I am grateful for the support of all the staff at IMR, but particularly Prescilla, Reidun, Helen and Sam, and none of this research would have been possible without the AQUASERV program. I am very grateful for the support I received and the people that I have met through this program and look forward for potential opportunities for collaboration in the future.